INTELIGENCE
https://medium.com/microbial-instincts/the-miraculous-origin-of-brain-intelligence-in-land-animals-like-us-5e288c356047
The Miraculous Origin of Brain Intelligence in Land Animals (Like Us)
When aquatic animals first transitioned to land, a virus-like element probably hijacked their neurons.
During the Devonian period, around 360–390 million years ago, sea animals took their first step onto the land, marking the first appearance of semiaquatic creatures called tetrapods.
Tetrapods had limbs with digits instead of fins, allowing them to move in shallow water and on land (Figure 1). They retained both lungs and gills, allowing them to breathe in both water and land. Over the next 60–90 million years, tetrapods evolved into amphibians and amniotes. Amniotes, in turn, diversified into reptiles, mammals, and birds, fully conquering terrestrial habitats over an additional 100–150 million years.
But during that venture onto land, an extraordinary event occurred: a virus-like element hijacked their germline cells. This invasion endowed their neurons with the capability to form synaptic connections vital for learning and memory. It was the beginning of land intelligence.
(Germline cells are the reproductive cells of an organism, such as sperm and eggs, which pass genetic information to the offspring. Germline genes are then present in every cell of the organism, including neurons.)

The ability of neurons to transfer genetic information and modulate synaptic connections through mechanisms similar to viruses underpins the brain’s capacity for learning, memory, and adaptation.
Arc 1: The Virus-like Behavior of Arc
Amidst the vast sea of academic literature — with about 4–5 million papers published annually — few discoveries stand out as true breakthroughs. One such breakthrough was a 2018 study in the prestigious journal Cell, conducted by scientists Pastuzyn et al. from the University of Utah’s Department of Neurobiology and Anatomy, USA.
Pastuzyn et al. studied the behavior and origin of the activity-regulated cytoskeleton-associated protein, shorthanded as Arc. This protein, first identified in the mammalian hippocampus in 1995, is known for its indispensable role in mediating neuroplasticity and memory formation. (The hippocampus is the brain region that stores memory.)
If the Arc gene is removed from mice, they become incapable of learning. For example, mice that find some cheese in a specific location won’t remember the route the following day.
Learning involves memory formation, which occurs when new synaptic connections are built and existing ones are strengthened between neurons in the brain — processes that require Arc. Synapses are the junctions where neurons meet one another. As follows, Arc inactivity has been associated with cognitive disorders and aging-related cognitive decline.
However, the mechanisms by which Arc mediates memory formation were unclear until Pastuzyn et al.’s groundbreaking study.
Pastuzyn et al. began by purifying some Arc proteins and visualizing them under the electron microscope. Surprisingly, Arc assembles into double-shell structures that remind you of viral capsids (Figure 2). Scientists—and even virologists, when shown these microscopy images—thought they were looking at viruses at first. But it wasn’t viruses.
Such protein purification techniques are highly precise. Standard practices like genetic sequencing and western blotting can confirm if the purified proteins’ genetic code or molecular structure matches the target, leaving no room for misattribution. So, they were undoubtedly looking at Arc.
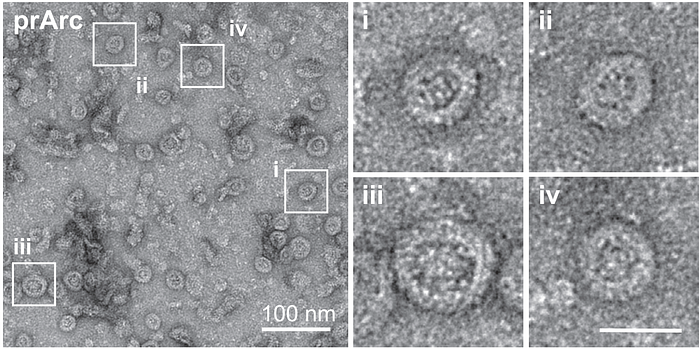
During the purification process, Pastuzyn et al. also observed that the Arc protein tends to co-purify with RNA, suggesting an interaction. Indeed, subsequent experiments revealed that Arc encapsulates messenger RNA (mRNA) within its shell, protecting it from degradation. This phenomenon mirrors how viruses carry genes inside their protective capsids.
Within a cell, mRNA carries the genetic instructions from the DNA in the nucleus to the ribosome, the site of protein synthesis. The ribosome then translates these genetic instructions into proteins. In essence, you can instruct a cell to make almost any protein if you can safely deliver the right mRNA gene sequence into the cell.
This is how Covid-19 mRNA vaccines instruct the cells to create spike proteins for immunization. This is how viruses instruct the cells they infect to create viral proteins and virions. And this is how Arc instructs the neurons to create new synaptic connections and memories.
Since Pastuyzn et al. already established that Arc encases mRNA inside its protective shell, they wanted to see how neurons use Arc. Through a series of sophisticated experiments, they showed that neurons release extracellular vesicles (EVs) that contain Arc.
Cells usually release EVs — which carry biological information such as proteins and genes — to relay signals to other cells. In this case, the neurons release Arc in their EVs to deliver mRNA to nearby neurons, instructing them to form new synaptic connections for memory formation.
“Here, we show that mammalian Arc protein exhibits many hallmarks of Gag proteins encoded by retroviruses and retrotransposons: self-assembly into capsids, RNA encapsulation, release in EVs, and intercellular transmission of RNA,” Pastuzyn et al. wrote in their paper.
“These data suggest that Arc can mediate intercellular trafficking of mRNA via Arc EVs, revealing a novel molecular mechanism by which genetic information may be transferred between neurons.”
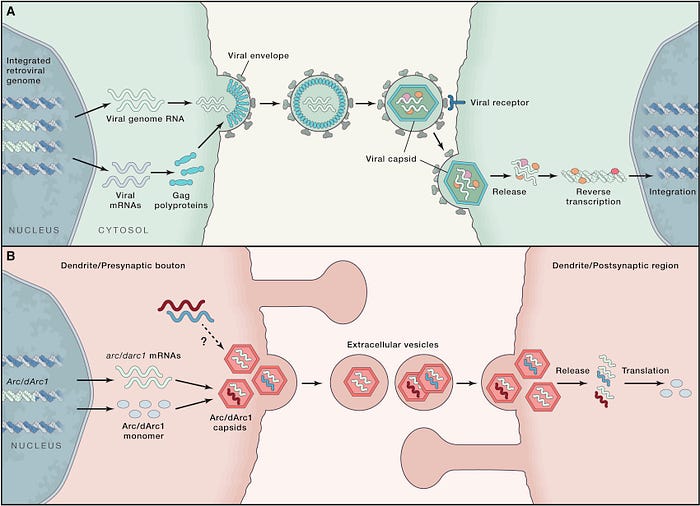
As Pastuzyn et al. summarized, the behavior of Arc in neurons is uncannily similar to that of viruses (Figure 3). Both form protective capsids that safeguard their genetic material. These capsids then deliver the genetic content to neighboring cells, influencing their cellular processes.
The virus-like behavior of Arc in hijacking and controlling nearby cells is unique to neurons. The ability of neurons to transfer genetic information and modulate synaptic connections through mechanisms similar to viruses underpins the brain’s capacity for learning, memory, and adaptation.
“I can see what people are thinking: Is memory a virus?” Jason Shepherd, PhD, laughed. Dr. Shepherd is an associate professor of neurobiology who directed the groundbreaking 2018 study by Pastuzyn et al.
Let’s have a brief recap of the key points so far:
- Arc is indispensable for learning and memory because it helps form new synaptic connections between neurons.
- In a groundbreaking discovery, Arc appears to behave like a virus. Arc forms capsids to encapsulate mRNA in extracellular vesicles (EVs), allowing the transfer of genetic information between neurons.
- This virus-like behavior of Arc is unique to neurons and is fundamental to the brain’s ability to learn and adapt by forming memories.
The next big question is: how did Arc evolve?
Arc didn’t exist before tetrapods, as the gene is absent in fish species — suggesting that Arc likely formed the dawn of land intelligence.
Arc 2: Origin of Arc
To decode evolutionary origins, scientists use phylogenetics to reconstruct the evolutionary historical tree of an organism or protein by comparing and backtracking its gene sequence across various species.
By aligning these sequences to map out a phylogenetic tree, scientists can trace the gene’s lineage back to a common ancestor and reveal how the gene has evolved and diverged over time.
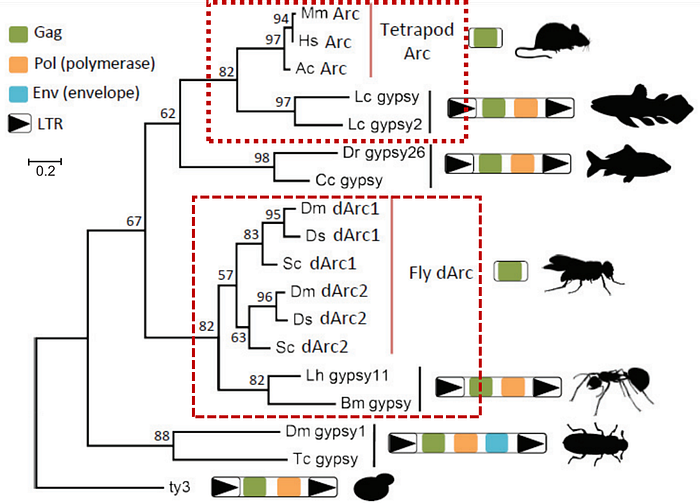
Pastuzyn et al. reconstructed the evolutionary tree of the Arc protein sequence, showing that Arc from mice (Mm, Mus musculus), humans (Hs, Homo sapiens), and lizard (As, Anolis carolinensis) matches that of the tetrapod — indicating that Arc is highly conserved and is largely unaltered since its emergence in tetrapods 360–390 million years ago.
Arc didn’t exist before tetrapods, as the gene is absent in fish species — suggesting that Arc mediated the beginnings of land intelligence.
The closest gene relative to tetrapod Arc is the Gag gene from gypsy elements of a fish species closely related to tetrapods (Lc, Latimeria chalumane) (see red annotation in Figure 4).
Interestingly, fruit flies (Dm, Drosophila melanogaster; Ds, Drosophila simulans; and Sc, Drosophila sechellia) also have the Arc protein, which shares evolutionary ties with the gypsy’s Gag gene from ants (Lh, Linepithema humile) and silkworm (Bm, Bombyx more).
Based on the evolutionary tree, tetrapod Arc and insect Arc have two different ancestors— indicating that the emergence of Arc occurred twice in land creatures (see red annotation in Figure 4).
Two or more independent evolution events of the same protein are not entirely unexpected. Convergent evolution occurs when different species independently evolve similar traits or functions, often in response to similar environmental pressures or functional needs.
Now, what’s Gag and gypsy, Arc’s closest evolutionary relative?
Gypsy is a retrotransposon, a self-replicating genetic element that can move around within an organism’s genome. Gypsy retrotransposons behave similarly to retroviruses by copying and inserting their genetic material into new locations within the DNA, which has contributed to genetic diversity throughout evolutionary history.
Retrotransposon isn’t a virus, although it’s very closely related to retroviruses — a class of viruses that includes human immunodeficiency virus (HIV). Both are characterized by their ability to insert their genetic material into the DNA of a host cell. Their main difference is that retroviruses can spread between organisms, while retrotransposons can only spread through reproduction to the offspring.
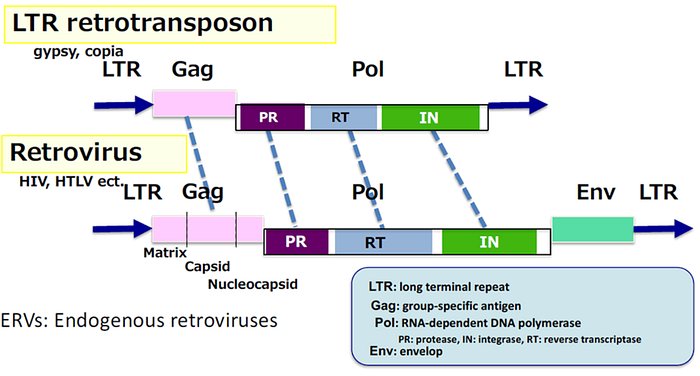
Regarding their genetic structure, retroviruses comprise the Gag, Pol, and Env genes. Retrotransposons only have the Gag and Pol genes (Figure 5). Gag encodes structural proteins that form capsids; Pol encodes enzymes necessary for integrating their genes into the host’s genome; and Env encodes envelope proteins needed for infectiousness. So, the only difference between them is the single piece of the Env gene.
As evident in their genetic structure, retroviruses and retrotransposons are so closely related that we don’t know which is the chicken or the egg. Retrotransposons could have either given rise to retroviruses (as an ancestor) or evolved from retroviruses (as a descendant). Or both of them could have descended from an unknown common ancestor.
Regardless, these retrotransposons and retroviruses form as much as 45–50% of the human genome, acquired through eons of evolution alongside ancient mobile genetic elements and viruses.
“Retrotransposons are considered either ancestral to, or derived from, modern retroviruses, but the line that distinguishes them is blurry,” explained a 2021 review article from Dr. Shepherd’s group. “Retrotransposons share sequence and structural similarities to bonafide retroviruses, and many contain virus-like genes.”
Moving on, Pastuzyn et al. found that Arc contains a sequence that corresponds to Gag. This aligns with the fact that the retrotransposon or retroviral Gag is responsible for encoding the structural protein needed for capsid formation. Yes, retrotransposons can form virus-like capsids to encapsulate their genetic material, protecting the genes for transfer and integration into new genomic locations like a retrovirus or Arc.
“Our data show a remarkable conservation of viral Gag properties in Arc,” Pastuzyn et al. wrote. “Since Arc shows structural homology to the Gag domain, the capability of self-assembly into oligomeric capsids is perhaps not too surprising.”
Based on this evidence, Pastuzyn et al. theorized that the retrotransposon Gag gene was likely domesticated or co-opted by early tetrapods to form Arc during their ventures onto the land. Why retrotransposon and not retrovirus, though? It’s because Arc’s closest evolutionary relative is the gypsy elements from fish, which are retrotransposons.
In other words, Arc’s sequence is most closely related to fish’s gypsy retrotransposons, and Arc’s sequence also contains a genetic domain that is the same as the retrotransposon/retroviral Gag gene.
Let’s recap briefly, as things may have gotten a bit complex.
- The closest evolutionary relative of Arc from humans and tetrapods is the Gag gene from gypsy retrotransposon elements of fish. Arc didn’t exist before tetrapods, as the gene is absent in fish species — suggesting that Arc likely formed the dawn of land intelligence.
- Gypsy is a retrotransposon (comprising Gag and Pol genes), a close evolutionary relative of retrovirus (comprising Gag, Pol, and Env genes). And it’s the retrotransposon/retroviral Gag gene that corresponds to Arc, suggesting that neurons may have domesticated Gag to form Arc.
- These retrotransposon and retrovirus sequences are already present in our genome and that of tetrapods, acquired via eons of evolution alongside ancient genetic elements and viruses.
It’s as if the cell intuitively knew how and when to evolve the Arc gene, unlocking the potential for higher-order cognition.
Arc 3: Evolution of Land Intelligence
Let’s move on to the final arc of this article — what actually happened when tetrapods ventured onto the land?
According to Pastuzyn et al.’s study, a Gag-containing retrotransposon — already present somewhere in the host genome — hijacked the germline cells and neurons of tetrapods. The neurons then repurposed this Gag gene in their genome to form Arc to help them learn how to learn — i.e., by forming memories — to survive in a foreign land.
“One explanation is that when animals moved onto land, there were all these new environments they had to adapt to — so you needed a smarter brain, and a more plastic brain,” Dr. Shepherd said. “Perhaps that is why Arc is so useful: It can give that extra plasticity or some sort of advantage.”
But how did the retrotransposon know which part of the genome to hijack? Or how did the cell know where to insert the retrotransposon sequence in its genome? One possible answer is that it doesn’t know.
“Sequencing of the human genome revealed that as much as 50% of the non-protein-coding sequences have viral or transposable element (TE) origins,” stated the 2021 review from Dr. Shepherd’s group. “In some cases, these random sequence insertions led to the generation of new genes with important functions.” (Italic emphasis mine).
If retrotransposon sequences are inserted into the host genome at random, then the domestication of the retrotransposon Gag gene into Arc in tetrapods’ neurons happened by chance — another testament to our luck. Our tetrapod ancestors somehow managed to evolve Arc to learn how to survive in a foreign land before they died without reproducing.
But what if there’s an alternative theory?
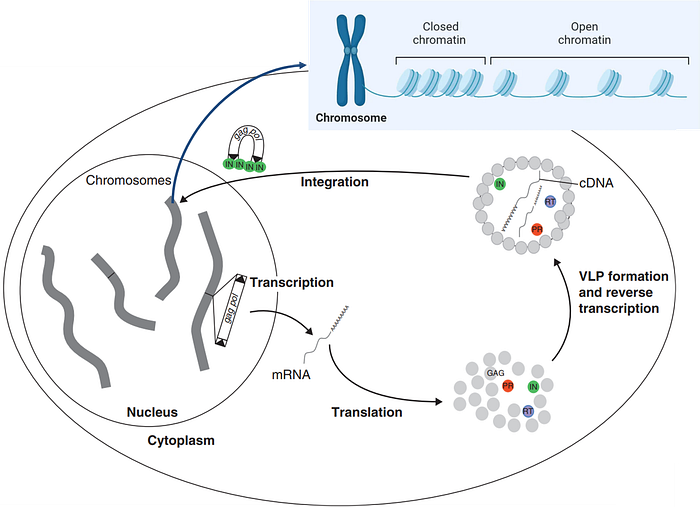
Some evidence indicates that retrotransposon integration isn’t random and prefers open chromatin regions (Figure 6). Chromatin is simply packaged DNA and proteins that form the chromosome in the cell nucleus; chromosomes then make up the cell’s genome. Open chromatin is the area where the DNA is loosely packed and more accessible for retrotransposon integration. Gene activities are also more active in these areas.
Thus, this selective integration into open chromatin regions (Figure 6) ensures that retrotransposons are maintained and propagated within the host genome, contributing to their survival and evolutionary success.
Other factors also influence how and where retrotransposon sequence gets integrated into the host genome. First, retrotransposons have integrase enzymes that recognize specific DNA sequences to be marked for integration. Second, the physical properties of DNA, such as flexibility and curvature, can also affect integration efficiency. Third, certain host proteins can guide retrotransposons to specific genomic sites. Fouth, epigenetic modifications like DNA methylation can influence chromatin accessibility and, thus, retrotransposon integration preference.
Basically, multiple factors can influence where the retrotransposon gets integrated into the host genome, making such events less random. This makes sense because if retrotransposon integration is truly random, our genomes would be so unstable and prone to harmful mutations.
If retrotransposon integration isn’t completely random, we’re back to the question of how the cell knew where to insert the retrotransposon sequence in its genome.
It’s hard to say for sure, but cells might be more intelligent than we think. They already had the building blocks to evolve Arc from a retrotransposon sequence within their genome. And, somehow, the cell managed to insert the retrotransposon Gag gene into its genome in a way that evolved into the Arc gene, and this happened just as tetrapods were transitioning onto land.
It’s as if the cell intuitively knew how and when to evolve the Arc gene, unlocking the potential for higher-order cognition.
How exactly did the cell manage this? We don’t know.
Evolutionary biologists often say that evolution doesn’t occur because cells or organisms ‘know’ what to do, but because natural selection favors certain evolutionary events and traits that provide an advantage in a given environment. This is what’s fascinating about evolutionary biology — the right changes happen at the right time and in the right place.
Finally, let’s have a recap of this final arc:
- The Gag-containing retrotransposon sequence — already present somewhere in the cell’s genome — somehow hijacked the germline cells and neurons of tetrapods. This event led to Arc evolution, enabling tetrapods to adapt to the new environmental pressures of terrestrial life by developing memory-forming capabilities.
- Retrotransposon integration isn’t entirely random. Some evidence shows retrotransposon prefers open chromatin regions, which are more accessible and genetically active. Other factors can also affect retrotransposon integration, making it less random than we think.
- But this doesn’t mean the cell or organism integrated the retrotransposon sequence with intention. Evolutionary biologists argue that evolution doesn’t occur because cells “know” what to do, but because natural selection favors certain evolutionary events and traits that provide an advantage in a given environment.
All in all, this is the miraculous story of how brain intelligence evolved in land dwellers like us. It’s the tale of the humble beginnings of higher-order cognitive processes that we should appreciate.
If you have made it this far, thank you for reading. Subscribe to my Medium email list here. You can also tip me here if you are feeling generous today, and I will appreciate any financial support I can get.


